Protein identification and mapping are cornerstones of proteomics, enabling researchers to unravel the structure, function, and dynamics of proteins in cells, tissues, and organisms. Understanding protein identity, modifications, and spatial distribution is crucial in fields like molecular biology, biomarker discovery, and systems biology.
🔍 What is Protein Identification?
Protein identification is the process of determining the amino acid sequence or name of a protein within a biological sample. It answers the fundamental question: What proteins are present in this sample?
Protein identification often follows protein separation, such as via SDS-PAGE or liquid chromatography, and utilizes analytical tools like mass spectrometry (MS) or Edman degradation to analyze the peptides or fragments.
🗺️ What is Protein Mapping?
Protein mapping involves determining the position, structure, and post-translational modifications (PTMs) of proteins. This includes:
- Peptide mapping: Breaking down proteins into peptides and analyzing their sequence and modifications.
- Epitope mapping: Identifying antibody-binding regions on proteins.
- 3D structural mapping: Using cryo-electron microscopy (Cryo-EM), X-ray crystallography, or NMR spectroscopy.
- Spatial mapping: Determining where proteins are located within a cell or tissue using imaging techniques.
🧪 Techniques for Protein Identification
1. Mass Spectrometry (MS)
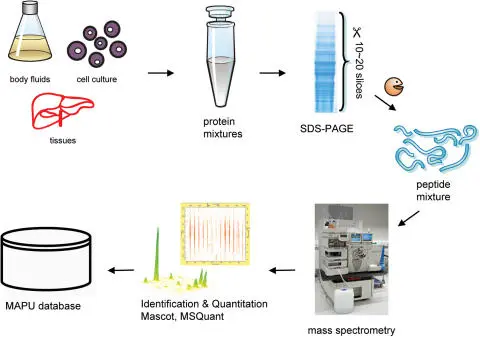
Mass spectrometry is the gold standard for protein identification. Key workflows include:
- Bottom-up proteomics: Proteins are digested into peptides (e.g., by trypsin), and the peptides are analyzed by MS.
- Top-down proteomics: Intact proteins are analyzed directly by MS, preserving PTMs.
Mass analyzers such as Orbitrap, TOF, or ion trap can detect peptide masses with high accuracy. MS data are compared against protein databases like UniProt or NCBI.
2. Edman Degradation
An older but precise chemical sequencing method that removes and identifies one amino acid at a time from the N-terminus of a protein. It’s limited to shorter sequences and requires a pure protein sample.
3. Western Blotting
This semi-quantitative method uses specific antibodies to detect known proteins after separation by SDS-PAGE and transfer to a membrane. It's useful for targeted identification but not for de novo sequencing.
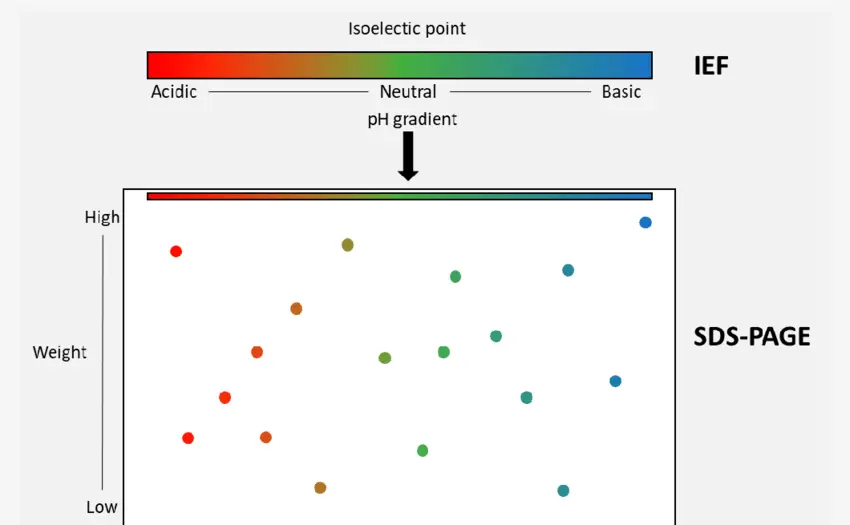
4. 2D Gel Electrophoresis (2D-PAGE)
Separates proteins by isoelectric point (pI) and molecular weight, allowing visualization of complex protein mixtures. Spots can be excised and identified by LC-MS/MS.
🧭 Techniques for Protein Mapping
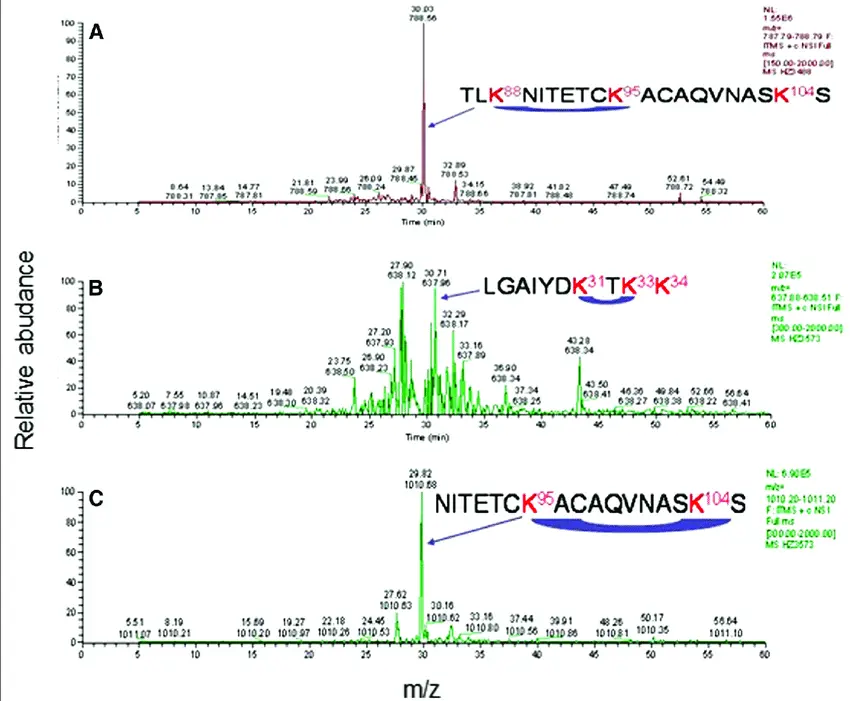
1. Peptide Mapping by LC-MS/MS
Peptide mapping uses enzymatic digestion followed by liquid chromatography tandem mass spectrometry (LC-MS/MS) to identify peptide sequences, PTMs (phosphorylation, glycosylation), and degradation products.
It’s widely used in:
- Biosimilar characterization
- Biopharmaceutical QC
- Enzyme activity studies
2. Epitope Mapping
Used in immunology and antibody development. Methods include:
- Peptide scanning
- X-ray crystallography of antigen-antibody complexes
- Hydrogen-deuterium exchange (HDX-MS)
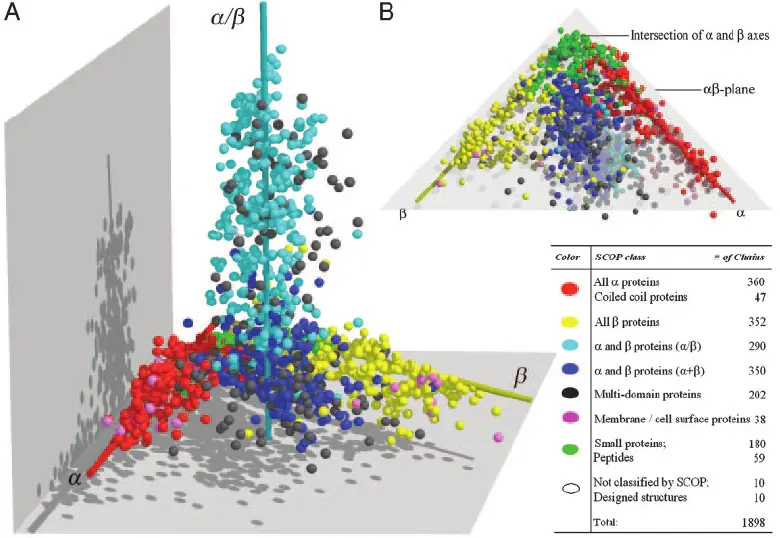
3. Structural Mapping (3D)
Determining atomic-level protein structures:
- X-ray crystallography: High-resolution but requires crystallization.
- Cryo-electron microscopy (Cryo-EM): Great for large complexes.
- Nuclear magnetic resonance (NMR): Ideal for smaller, soluble proteins.
4. Immunofluorescence and Imaging
Localizes proteins in tissues or cells using:
- Confocal microscopy
- Immunohistochemistry (IHC)
- Mass spectrometry imaging (MSI)
🔬 Applications of Protein Identification and Mapping
✅ Biomedical Research
- Identifying disease biomarkers (e.g., Alzheimer’s, cancer)
- Analyzing protein networks in health vs. disease
- Understanding mechanisms of drug action
✅ Diagnostics
- Identifying autoantigens in autoimmune diseases
- Discovering microbial proteins in infectious diseases
- Creating targeted immunoassays
✅ Personalized Medicine
- Profiling patient-specific proteomes
- Predicting drug response or resistance
- Monitoring disease progression via protein biomarkers
🚀 Advances and Future Trends
1. Single-Cell Proteomics
Emerging technologies now allow proteomic profiling at the single-cell level, enabling the study of cellular heterogeneity in tumors and tissues.
2. AI and Machine Learning
Machine learning enhances peptide matching, PTM prediction, and structural modeling (AlphaFold 3 for protein folding prediction).
3. Label-Free Quantification
Advancements in label-free MS quantitation improve dynamic proteome profiling without the need for isotopic labels.
4. Spatial Omics
Combining proteomics with spatial transcriptomics and metabolomics to create comprehensive tissue maps.
Conclusion
Protein identification and mapping are essential for decoding the complexity of biological systems. Whether you're studying disease mechanisms, developing therapeutics, or building diagnostic assays, understanding protein profiles at both sequence and structural levels is key.
With ever-improving technologies like high resolution MS, AIpowered protein modeling, and single cell proteomics, the field continues to evolve offering unprecedented insight into the dynamic proteome.